This article provides multiple solutions to create an elegant Venn diagram with R or RStudio. The following R packages will be illustrated: ggvenn, ggVenDiagram, VennDiagram and the gplots packages.
Contents:
Create a demo data
set.seed(20190708)
genes <- paste("gene",1:1000,sep="")
x <- list(
A = sample(genes,300),
B = sample(genes,525),
C = sample(genes,440),
D = sample(genes,350)
)Using the ggvenn R package
This package is a ggplot2 extension. A color can be assigned to each set.
Installation:
if (!require(devtools)) install.packages("devtools")
devtools::install_github("yanlinlin82/ggvenn")Usage:
library(ggvenn)
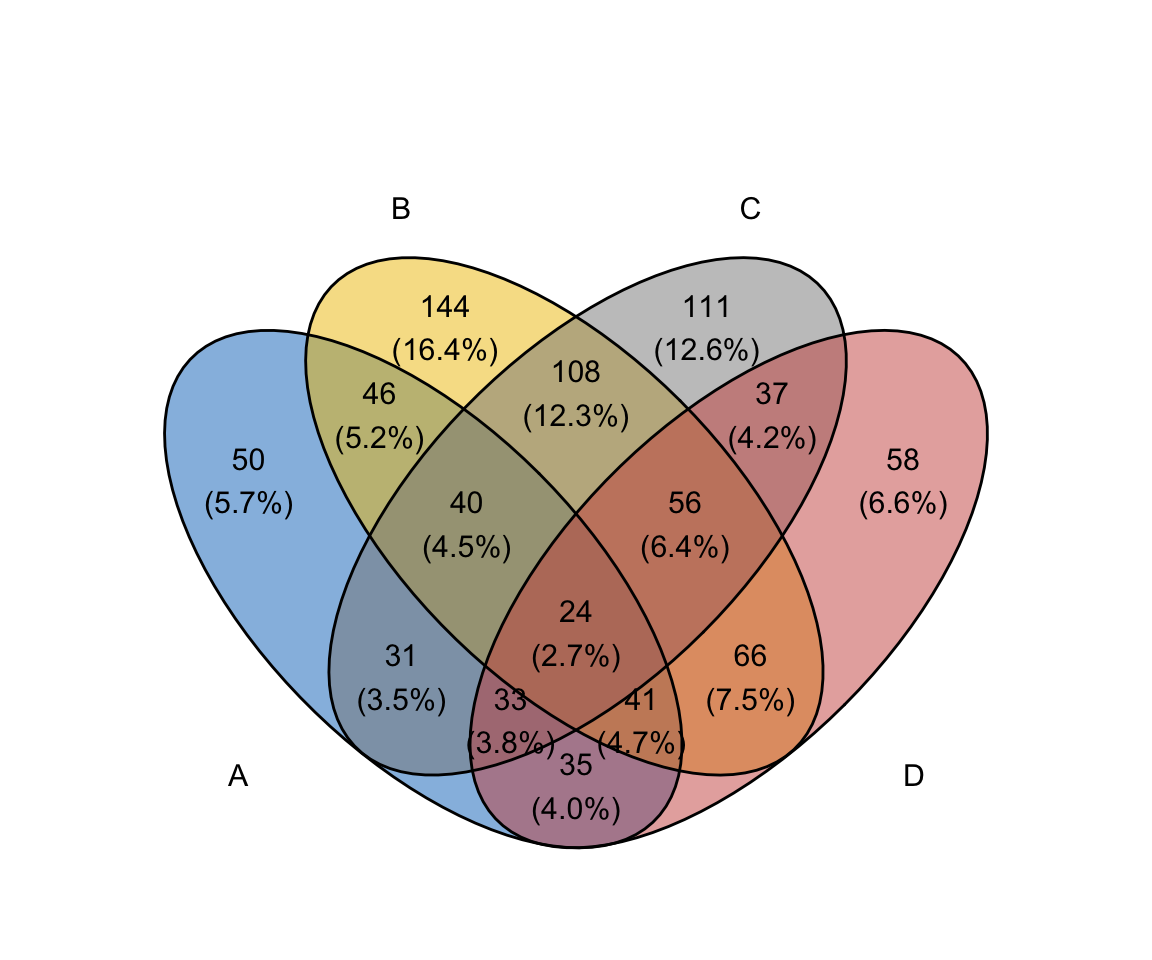
ggvenn(
x,
fill_color = c("#0073C2FF", "#EFC000FF", "#868686FF", "#CD534CFF"),
stroke_size = 0.5, set_name_size = 4
)
Read more: Beautiful GGPlot Venn Diagram with R
Using the ggVennDiagram R package
This package is a ggplot2 extension. The ggVennDiagram package maps the fill color of each region to quantity, allowing us to visually observe the differences between different parts.
Installation:
if (!require(devtools)) install.packages("devtools")
devtools::install_github("gaospecial/ggVennDiagram")Usage:
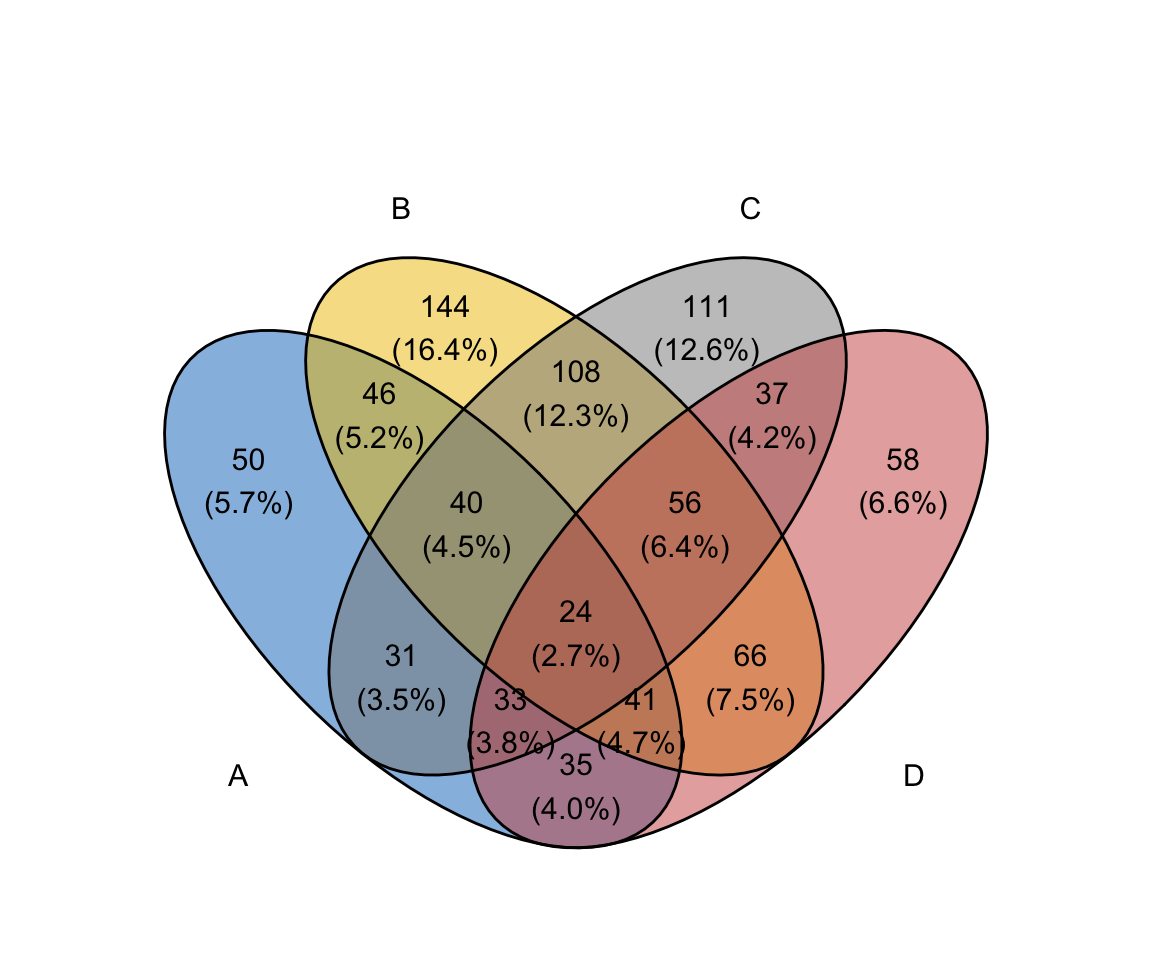
library("ggVennDiagram")
ggVennDiagram(x, label_alpha = 0)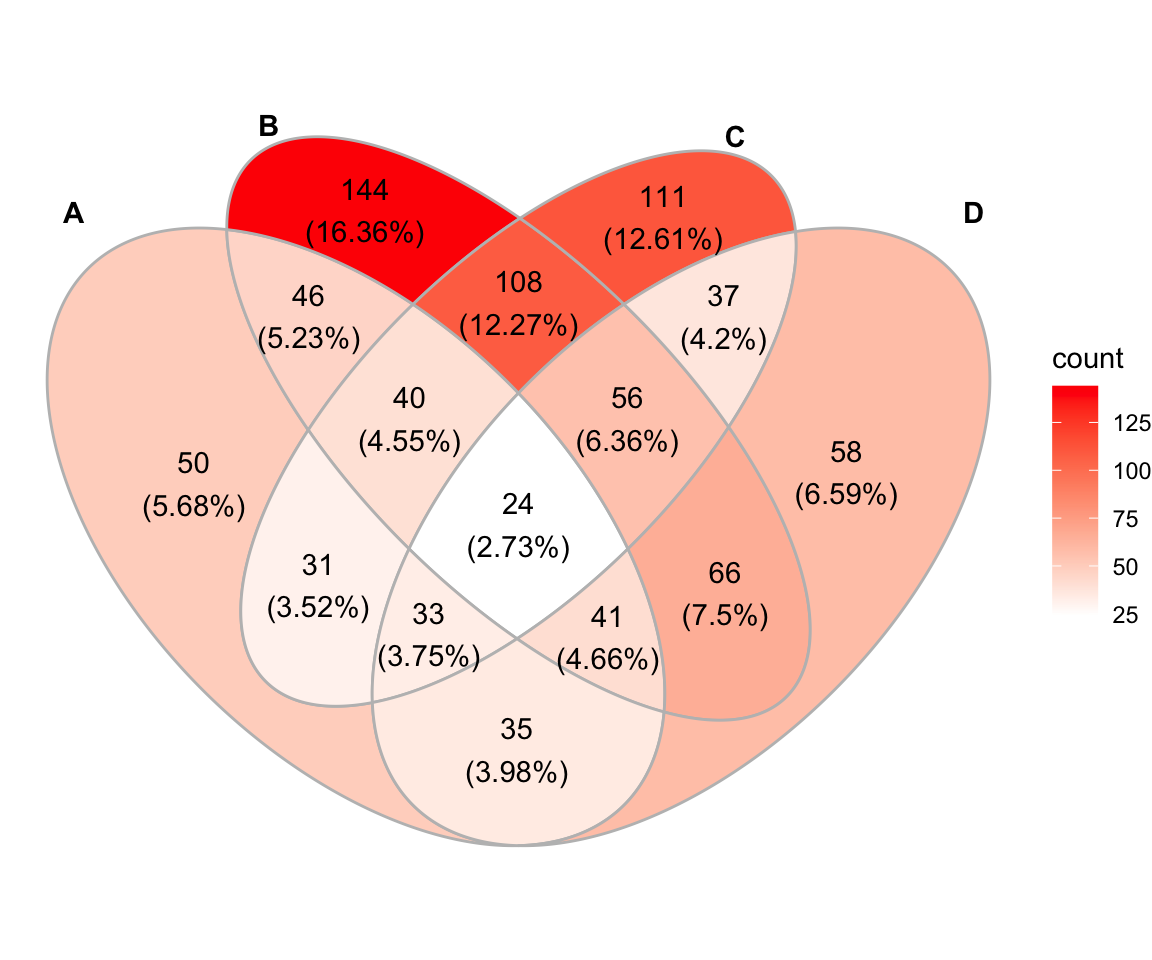
Read more: Beautiful GGPlot Venn Diagram with R
Using the VennDiagram R package
Installation:
install.packages("VennDiagram")`Usage:
Create a Venn diagram and save it into a file. The function venn.diagram() takes a list and creates a file containing a publication-quality Venn Diagram.
library(VennDiagram)
venn.diagram(x, filename = "venn-4-dimensions.png")Display the plot directly in R:
# Helper function to display Venn diagram
display_venn <- function(x, ...){
library(VennDiagram)
grid.newpage()
venn_object <- venn.diagram(x, filename = NULL, ...)
grid.draw(venn_object)
}# Four dimension Venn plot
display_venn(x)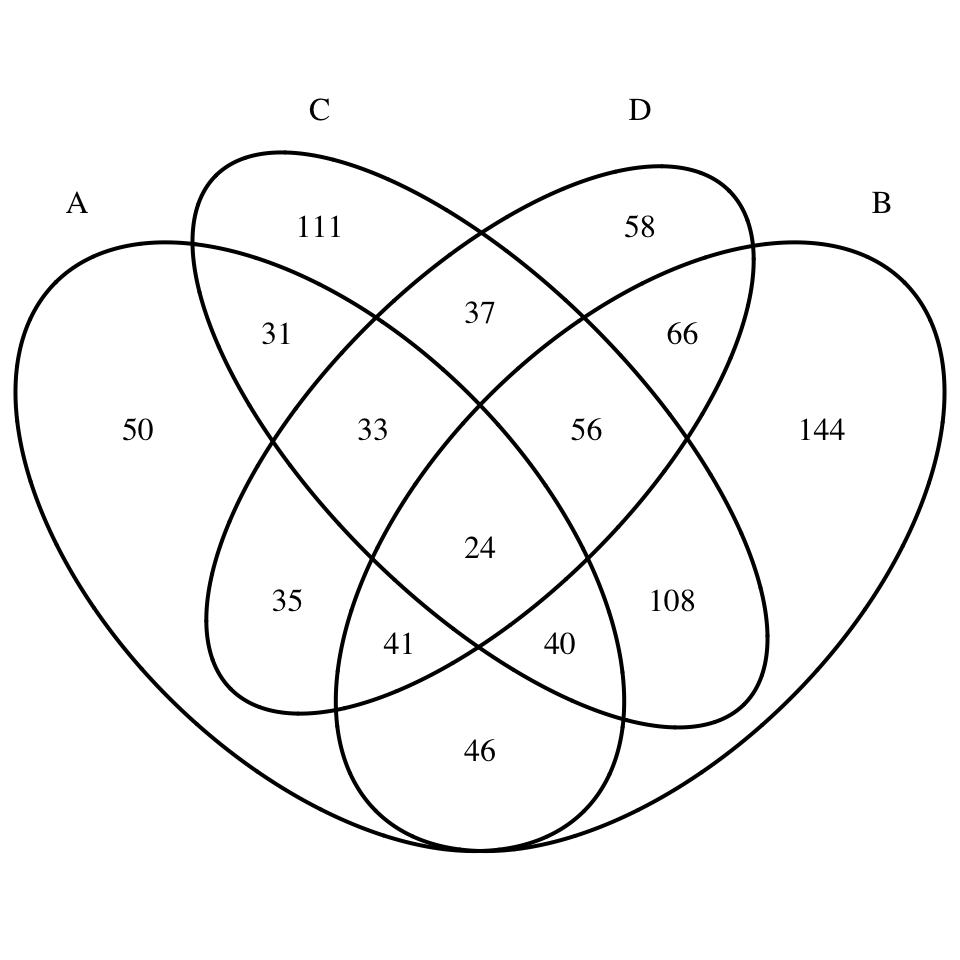
# Three dimension Venn plot
display_venn(x[1:3])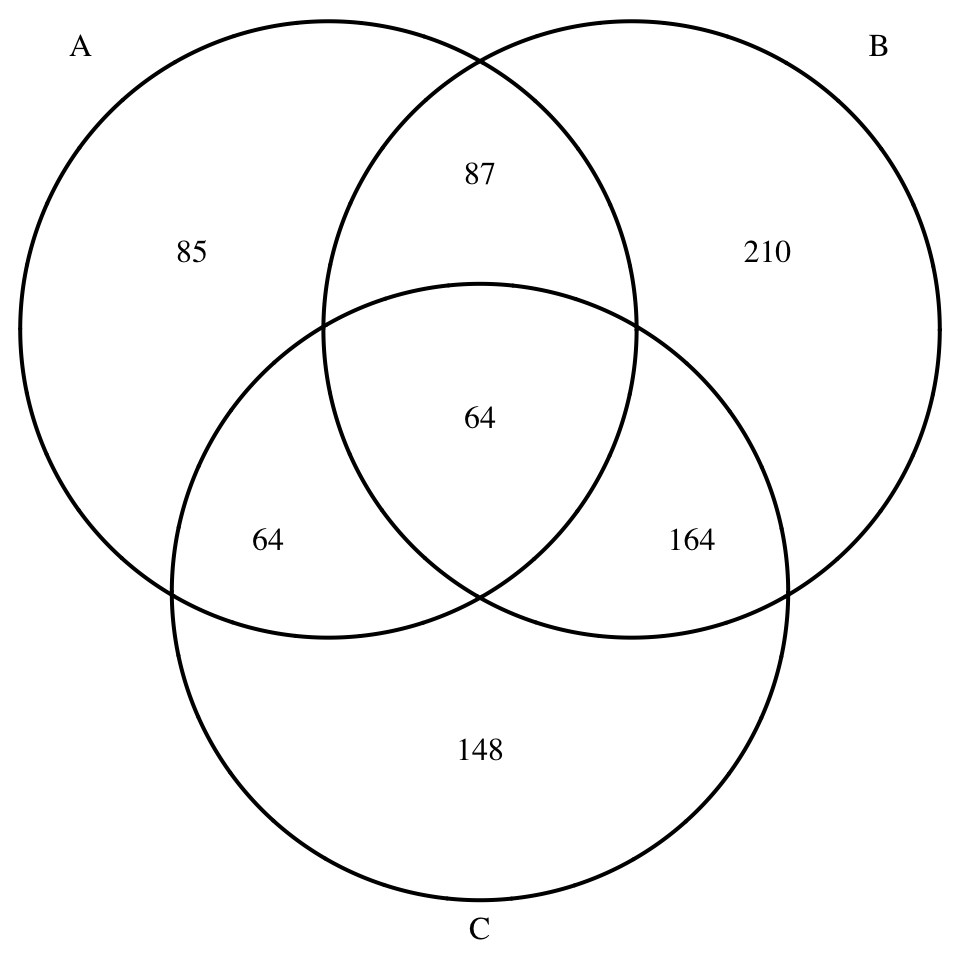
# Change category names
# Change fill color
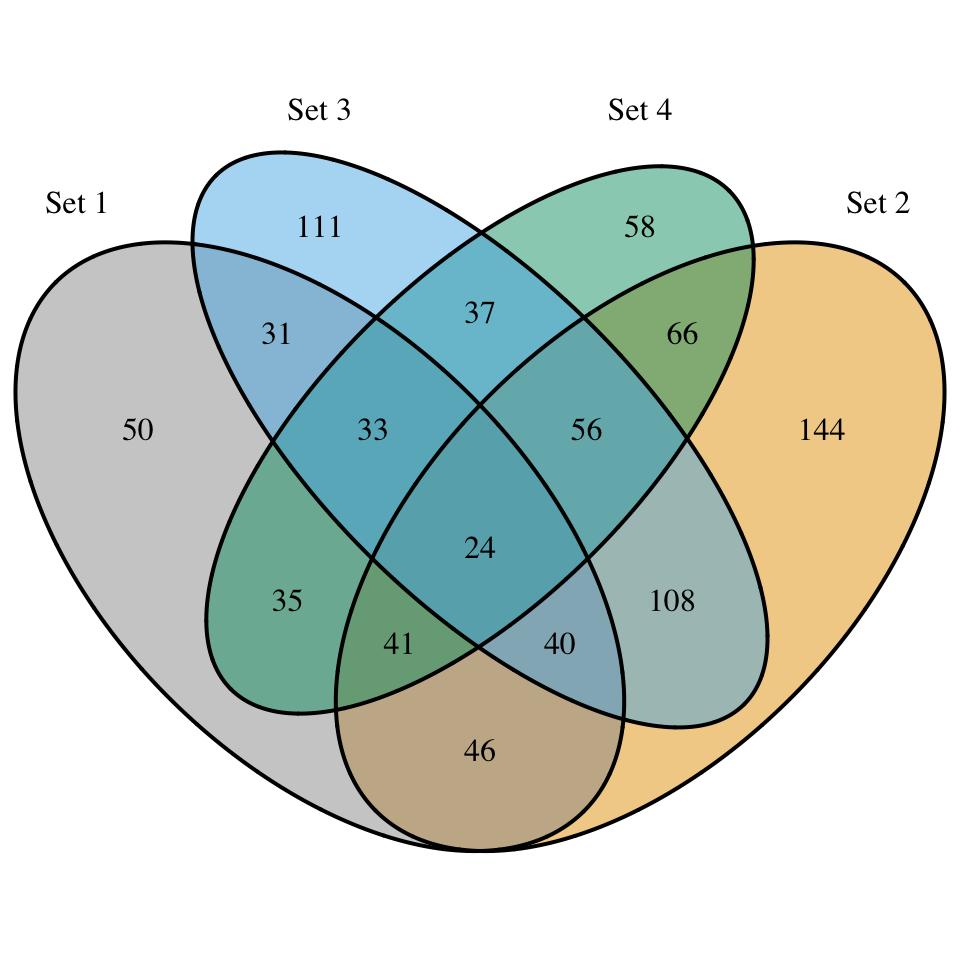
display_venn(
x,
category.names = c("Set 1" , "Set 2 " , "Set 3", "Set 4"),
fill = c("#999999", "#E69F00", "#56B4E9", "#009E73")
)
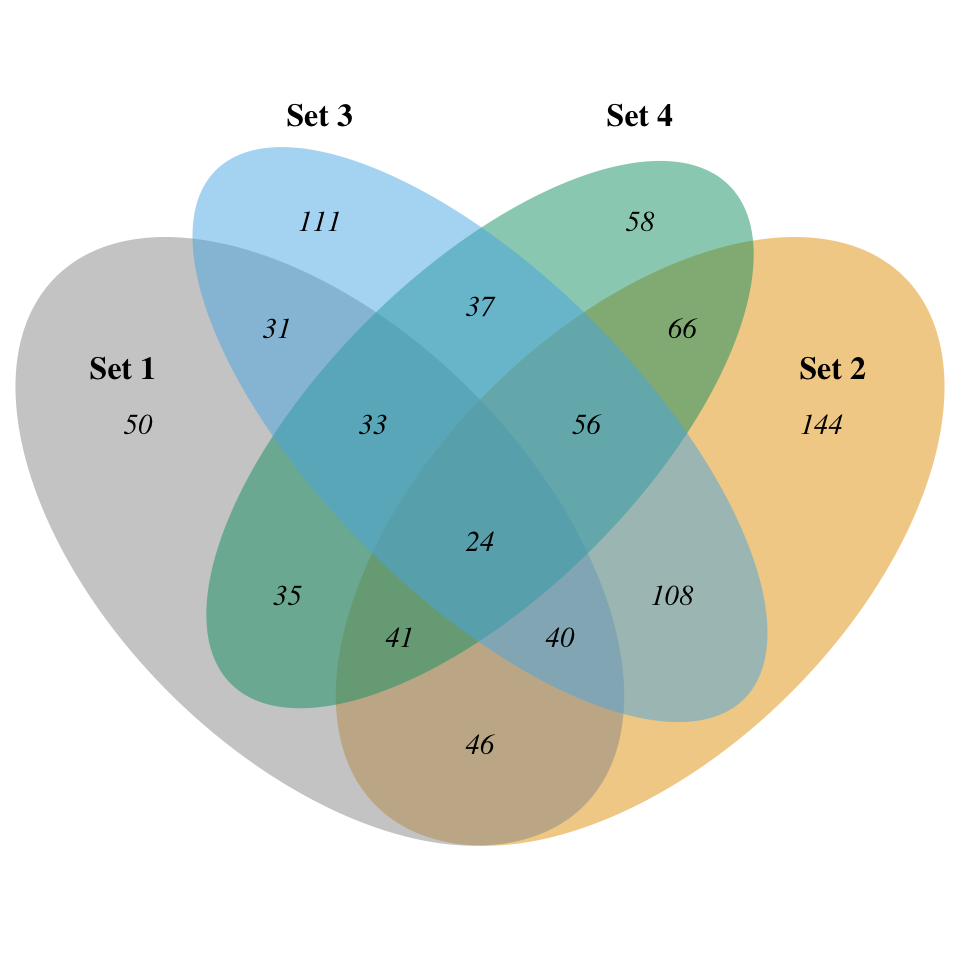
# Further customization
display_venn(
x,
category.names = c("Set 1" , "Set 2 " , "Set 3", "Set 4"),
# Circles
lwd = 2,
lty = 'blank',
fill = c("#999999", "#E69F00", "#56B4E9", "#009E73"),
# Numbers
cex = .9,
fontface = "italic",
# Set names
cat.cex = 1,
cat.fontface = "bold",
cat.default.pos = "outer",
cat.dist = c(0.055, 0.055, 0.1, 0.1)
)
Using the gplots R package
Installation:
install.package("gplots")`Usage:
library(gplots)
v.table <- venn(x)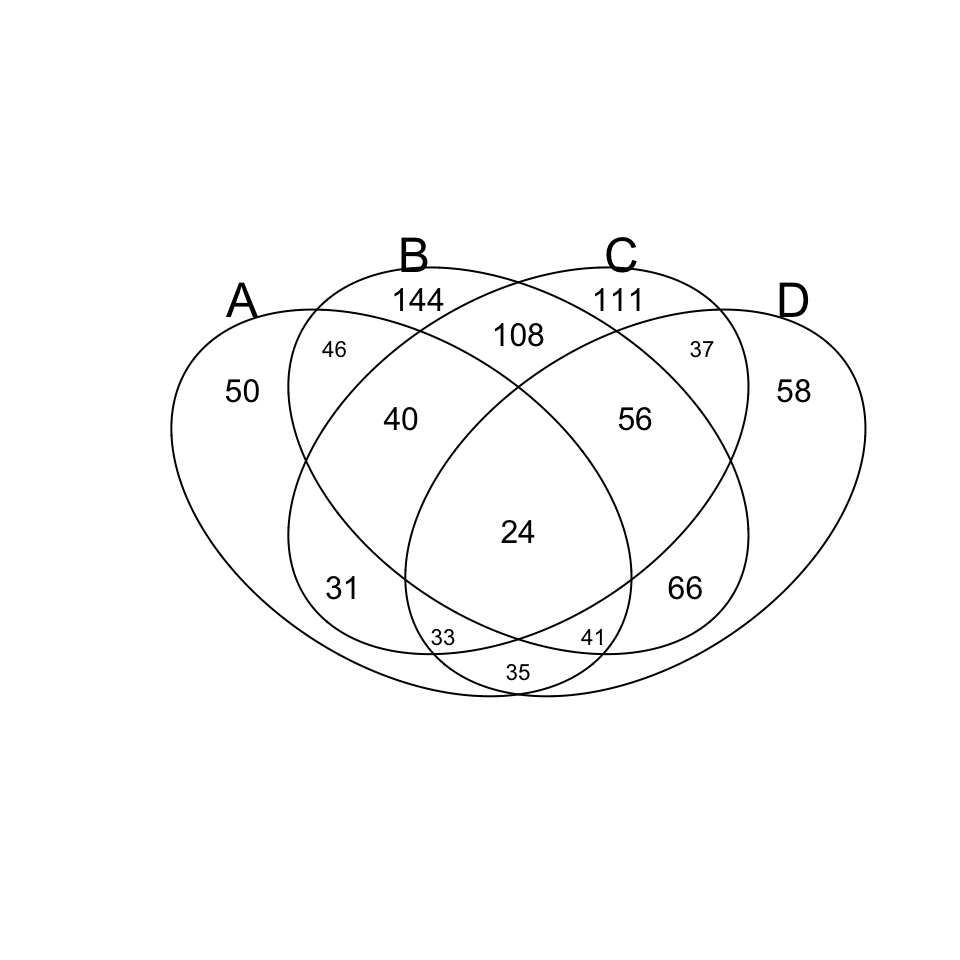
Access to the intersection data:
print(v.table)## num A B C D
## 0000 0 0 0 0 0
## 0001 58 0 0 0 1
## 0010 111 0 0 1 0
## 0011 37 0 0 1 1
## 0100 144 0 1 0 0
## 0101 66 0 1 0 1
## 0110 108 0 1 1 0
## 0111 56 0 1 1 1
## 1000 50 1 0 0 0
## 1001 35 1 0 0 1
## 1010 31 1 0 1 0
## 1011 33 1 0 1 1
## 1100 46 1 1 0 0
## 1101 41 1 1 0 1
## 1110 40 1 1 1 0
## 1111 24 1 1 1 1
## attr(,"intersections")
## attr(,"intersections")$A
## [1] "gene835" "gene910" "gene866" "gene785" "gene532" "gene385" "gene201" "gene74" "gene941" "gene401" "gene642"
## [12] "gene299" "gene819" "gene245" "gene834" "gene700" "gene422" "gene176" "gene643" "gene852" "gene584" "gene560"
## [23] "gene638" "gene164" "gene122" "gene419" "gene521" "gene363" "gene724" "gene160" "gene988" "gene328" "gene452"
## [34] "gene236" "gene707" "gene865" "gene342" "gene165" "gene572" "gene565" "gene206" "gene663" "gene107" "gene620"
## [45] "gene917" "gene298" "gene402" "gene840" "gene992" "gene225"
##
## attr(,"intersections")$B
## [1] "gene129" "gene150" "gene307" "gene73" "gene651" "gene40" "gene722" "gene374" "gene779" "gene772"
## [11] "gene574" "gene355" "gene970" "gene429" "gene979" "gene656" "gene208" "gene274" "gene252" "gene585"
## [21] "gene662" "gene535" "gene709" "gene589" "gene964" "gene569" "gene113" "gene851" "gene111" "gene308"
## [31] "gene18" "gene309" "gene977" "gene530" "gene929" "gene720" "gene83" "gene256" "gene217" "gene287"
## [41] "gene596" "gene776" "gene573" "gene99" "gene470" "gene689" "gene44" "gene473" "gene380" "gene246"
## [51] "gene774" "gene799" "gene621" "gene134" "gene659" "gene947" "gene322" "gene397" "gene664" "gene266"
## [61] "gene556" "gene210" "gene272" "gene389" "gene84" "gene771" "gene55" "gene349" "gene51" "gene137"
## [71] "gene462" "gene106" "gene353" "gene801" "gene943" "gene297" "gene678" "gene546" "gene617" "gene289"
## [81] "gene480" "gene361" "gene913" "gene875" "gene948" "gene777" "gene788" "gene743" "gene199" "gene879"
## [91] "gene229" "gene951" "gene447" "gene175" "gene715" "gene942" "gene668" "gene698" "gene559" "gene812"
## [101] "gene52" "gene337" "gene413" "gene757" "gene472" "gene526" "gene876" "gene567" "gene813" "gene147"
## [111] "gene685" "gene114" "gene614" "gene427" "gene508" "gene247" "gene540" "gene986" "gene241" "gene731"
## [121] "gene591" "gene279" "gene120" "gene346" "gene403" "gene333" "gene922" "gene135" "gene200" "gene603"
## [131] "gene914" "gene820" "gene492" "gene19" "gene978" "gene804" "gene655" "gene332" "gene958" "gene903"
## [141] "gene564" "gene989" "gene386" "gene151"
##
## attr(,"intersections")$C
## [1] "gene169" "gene390" "gene265" "gene581" "gene250" "gene634" "gene188" "gene319" "gene994" "gene366"
## [11] "gene213" "gene610" "gene661" "gene654" "gene453" "gene935" "gene193" "gene251" "gene243" "gene545"
## [21] "gene178" "gene411" "gene737" "gene953" "gene902" "gene557" "gene691" "gene221" "gene920" "gene495"
## [31] "gene474" "gene543" "gene695" "gene100" "gene686" "gene383" "gene125" "gene608" "gene278" "gene59"
## [41] "gene895" "gene828" "gene288" "gene611" "gene441" "gene396" "gene93" "gene448" "gene428" "gene367"
## [51] "gene438" "gene459" "gene579" "gene395" "gene248" "gene867" "gene45" "gene29" "gene437" "gene77"
## [61] "gene70" "gene484" "gene633" "gene85" "gene854" "gene490" "gene177" "gene426" "gene887" "gene550"
## [71] "gene923" "gene792" "gene412" "gene212" "gene590" "gene292" "gene795" "gene30" "gene159" "gene304"
## [81] "gene471" "gene62" "gene790" "gene28" "gene770" "gene533" "gene575" "gene173" "gene832" "gene860"
## [91] "gene683" "gene878" "gene818" "gene944" "gene789" "gene357" "gene391" "gene888" "gene919" "gene796"
## [101] "gene631" "gene82" "gene733" "gene732" "gene197" "gene368" "gene909" "gene816" "gene843" "gene765"
## [111] "gene898"
##
## attr(,"intersections")$D
## [1] "gene738" "gene321" "gene149" "gene938" "gene487" "gene145" "gene783" "gene15" "gene871" "gene475" "gene528"
## [12] "gene706" "gene130" "gene570" "gene43" "gene489" "gene640" "gene161" "gene293" "gene966" "gene171" "gene498"
## [23] "gene926" "gene701" "gene632" "gene649" "gene853" "gene702" "gene869" "gene831" "gene392" "gene694" "gene271"
## [34] "gene650" "gene880" "gene906" "gene541" "gene538" "gene420" "gene140" "gene677" "gene124" "gene644" "gene233"
## [45] "gene868" "gene152" "gene800" "gene12" "gene496" "gene39" "gene726" "gene674" "gene226" "gene379" "gene916"
## [56] "gene602" "gene857" "gene523"
##
## attr(,"intersections")$`A:B`
## [1] "gene168" "gene301" "gene86" "gene133" "gene786" "gene235" "gene209" "gene205" "gene451" "gene372" "gene794"
## [12] "gene889" "gene838" "gene604" "gene280" "gene848" "gene249" "gene965" "gene877" "gene599" "gene263" "gene607"
## [23] "gene998" "gene17" "gene561" "gene592" "gene123" "gene186" "gene270" "gene872" "gene96" "gene170" "gene645"
## [34] "gene817" "gene239" "gene957" "gene993" "gene222" "gene154" "gene510" "gene983" "gene476" "gene658" "gene253"
## [45] "gene773" "gene973"
##
## attr(,"intersections")$`A:C`
## [1] "gene393" "gene20" "gene71" "gene479" "gene343" "gene826" "gene56" "gene87" "gene8" "gene50" "gene115"
## [12] "gene464" "gene981" "gene760" "gene485" "gene54" "gene255" "gene822" "gene892" "gene359" "gene325" "gene117"
## [23] "gene769" "gene805" "gene497" "gene516" "gene855" "gene488" "gene553" "gene763" "gene862"
##
## attr(,"intersections")$`A:D`
## [1] "gene982" "gene46" "gene901" "gene697" "gene782" "gene635" "gene563" "gene692" "gene269" "gene657" "gene132"
## [12] "gene25" "gene261" "gene211" "gene486" "gene710" "gene991" "gene126" "gene870" "gene121" "gene404" "gene499"
## [23] "gene945" "gene833" "gene172" "gene219" "gene830" "gene406" "gene345" "gene797" "gene444" "gene3" "gene64"
## [34] "gene544" "gene10"
##
## attr(,"intersections")$`B:C`
## [1] "gene703" "gene874" "gene27" "gene627" "gene182" "gene254" "gene41" "gene163" "gene967"
## [10] "gene257" "gene975" "gene899" "gene962" "gene494" "gene845" "gene189" "gene47" "gene326"
## [19] "gene109" "gene940" "gene514" "gene334" "gene588" "gene897" "gene780" "gene264" "gene158"
## [28] "gene424" "gene600" "gene750" "gene587" "gene858" "gene766" "gene511" "gene524" "gene500"
## [37] "gene856" "gene515" "gene861" "gene320" "gene240" "gene338" "gene939" "gene849" "gene810"
## [46] "gene423" "gene775" "gene69" "gene752" "gene622" "gene502" "gene713" "gene542" "gene755"
## [55] "gene431" "gene153" "gene905" "gene598" "gene667" "gene439" "gene192" "gene88" "gene764"
## [64] "gene194" "gene408" "gene282" "gene884" "gene626" "gene162" "gene90" "gene335" "gene995"
## [73] "gene823" "gene273" "gene295" "gene183" "gene468" "gene315" "gene1000" "gene507" "gene348"
## [82] "gene259" "gene803" "gene911" "gene577" "gene1" "gene728" "gene873" "gene223" "gene101"
## [91] "gene63" "gene781" "gene446" "gene864" "gene81" "gene196" "gene195" "gene615" "gene354"
## [100] "gene719" "gene954" "gene549" "gene23" "gene376" "gene613" "gene809" "gene606" "gene26"
##
## attr(,"intersections")$`B:D`
## [1] "gene456" "gene747" "gene839" "gene699" "gene759" "gene364" "gene230" "gene327" "gene886" "gene268" "gene14"
## [12] "gene527" "gene260" "gene112" "gene985" "gene204" "gene681" "gene595" "gene36" "gene128" "gene824" "gene180"
## [23] "gene352" "gene351" "gene971" "gene660" "gene7" "gene224" "gene744" "gene609" "gene793" "gene666" "gene97"
## [34] "gene136" "gene506" "gene285" "gene156" "gene946" "gene725" "gene102" "gene883" "gene33" "gene387" "gene605"
## [45] "gene740" "gene339" "gene329" "gene139" "gene568" "gene432" "gene980" "gene504" "gene505" "gene517" "gene460"
## [56] "gene687" "gene629" "gene218" "gene671" "gene734" "gene375" "gene932" "gene167" "gene639" "gene407" "gene416"
##
## attr(,"intersections")$`C:D`
## [1] "gene399" "gene960" "gene531" "gene708" "gene323" "gene837" "gene704" "gene381" "gene95" "gene458" "gene503"
## [12] "gene493" "gene443" "gene915" "gene281" "gene891" "gene716" "gene636" "gene220" "gene762" "gene179" "gene696"
## [23] "gene844" "gene42" "gene314" "gene184" "gene394" "gene937" "gene896" "gene672" "gene918" "gene103" "gene324"
## [34] "gene721" "gene286" "gene141" "gene928"
##
## attr(,"intersections")$`A:B:C`
## [1] "gene882" "gene157" "gene49" "gene119" "gene384" "gene5" "gene166" "gene807" "gene586" "gene296" "gene969"
## [12] "gene231" "gene108" "gene317" "gene227" "gene934" "gene554" "gene421" "gene316" "gene418" "gene16" "gene330"
## [23] "gene481" "gene714" "gene53" "gene340" "gene730" "gene347" "gene283" "gene181" "gene727" "gene571" "gene836"
## [34] "gene679" "gene445" "gene501" "gene933" "gene410" "gene529" "gene94"
##
## attr(,"intersections")$`A:B:D`
## [1] "gene904" "gene79" "gene207" "gene382" "gene331" "gene753" "gene924" "gene356" "gene142" "gene463" "gene341"
## [12] "gene537" "gene829" "gene144" "gene21" "gene815" "gene4" "gene729" "gene646" "gene931" "gene997" "gene312"
## [23] "gene859" "gene594" "gene465" "gene684" "gene949" "gene539" "gene440" "gene398" "gene425" "gene174" "gene597"
## [34] "gene405" "gene653" "gene612" "gene75" "gene927" "gene378" "gene370" "gene908"
##
## attr(,"intersections")$`A:C:D`
## [1] "gene449" "gene369" "gene78" "gene414" "gene358" "gene749" "gene984" "gene893" "gene37" "gene32" "gene148"
## [12] "gene718" "gene907" "gene138" "gene739" "gene723" "gene578" "gene930" "gene336" "gene680" "gene234" "gene455"
## [23] "gene547" "gene756" "gene388" "gene688" "gene717" "gene754" "gene758" "gene61" "gene552" "gene518" "gene202"
##
## attr(,"intersections")$`B:C:D`
## [1] "gene558" "gene131" "gene238" "gene65" "gene536" "gene955" "gene630" "gene6" "gene641" "gene682" "gene802"
## [12] "gene576" "gene900" "gene827" "gene814" "gene433" "gene624" "gene601" "gene127" "gene228" "gene310" "gene652"
## [23] "gene670" "gene841" "gene894" "gene936" "gene365" "gene711" "gene616" "gene974" "gene566" "gene216" "gene921"
## [34] "gene466" "gene2" "gene302" "gene311" "gene806" "gene520" "gene811" "gene191" "gene277" "gene360" "gene990"
## [45] "gene232" "gene469" "gene400" "gene214" "gene665" "gene72" "gene846" "gene31" "gene92" "gene583" "gene242"
## [56] "gene483"
##
## attr(,"intersections")$`A:B:C:D`
## [1] "gene371" "gene735" "gene787" "gene38" "gene66" "gene415" "gene284" "gene477" "gene768" "gene454" "gene522"
## [12] "gene784" "gene582" "gene34" "gene618" "gene741" "gene850" "gene373" "gene742" "gene912" "gene748" "gene291"
## [23] "gene863" "gene143"
##
## attr(,"class")
## [1] "venn"Conclusion
This article describes multiple solutions to create Venn diagrams in R.
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Coursera - Online Courses and Specialization
Data science
- Course: Machine Learning: Master the Fundamentals by Stanford
- Specialization: Data Science by Johns Hopkins University
- Specialization: Python for Everybody by University of Michigan
- Courses: Build Skills for a Top Job in any Industry by Coursera
- Specialization: Master Machine Learning Fundamentals by University of Washington
- Specialization: Statistics with R by Duke University
- Specialization: Software Development in R by Johns Hopkins University
- Specialization: Genomic Data Science by Johns Hopkins University
Popular Courses Launched in 2020
- Google IT Automation with Python by Google
- AI for Medicine by deeplearning.ai
- Epidemiology in Public Health Practice by Johns Hopkins University
- AWS Fundamentals by Amazon Web Services
Trending Courses
- The Science of Well-Being by Yale University
- Google IT Support Professional by Google
- Python for Everybody by University of Michigan
- IBM Data Science Professional Certificate by IBM
- Business Foundations by University of Pennsylvania
- Introduction to Psychology by Yale University
- Excel Skills for Business by Macquarie University
- Psychological First Aid by Johns Hopkins University
- Graphic Design by Cal Arts
Amazon FBA
Amazing Selling Machine
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Version:
 Français
Français






Hi i am Venkata Panchumarthi.
Awesome. This article is very nice and it gives lot of information. I read the excellent article in recent days its helped me a lot. Thanks for sharing such a nice article.