In biomedical, behavioral research and many other fields, it is frequently required that a group of participants is rated or classified into categories by two observers (or raters, methods, etc). An example is two clinicians that classify the extent of disease in patients. The analysis of the agreement between the two observers can be used to measure the reliability of the rating system. High agreement would indicate consensus in the diagnosis and interchangeability of the observers (Warrens 2013).
In a previous chapter (Chapter @ref(cohen-s-kappa)), we described the classical Cohen’s Kappa, which is a popular measure of inter-rater reliability or inter-rater agreement. The Classical Cohen’s Kappa only counts strict agreement, where the same category is assigned by both raters (Friendly, Meyer, and Zeileis 2015). It takes no account of the degree of disagreement, all disagreements are treated equally. This is most appropriate when you have nominal variables. For ordinal rating scale it may preferable to give different weights to the disagreements depending on the magnitude.
This chapter describes the weighted kappa, a variant of the Cohen’s Kappa, that allows partial agreement (J. Cohen 1968). In other words, the weighted kappa allows the use of weighting schemes to take into account the closeness of agreement between categories. This is only suitable in the situation where you have ordinal or ranked variables.
Recall that, the kappa coefficients remove the chance agreement, which is the proportion of agreement that you would expect two raters to have based simply on chance.
Here, you will learn:
- Basics and the formula of the weighted kappa
- Assumptions and requirements for computing the weighted kappa
- Examples of R code for computing the weighted kappa
Contents:
Related Book
Inter-Rater Reliability Essentials: Practical Guide in RPrerequisites
Read the Chapter on Cohen’s Kappa (Chapter @ref(cohen-s-kappa)).
Basics
To explain the basic concept of the weighted kappa, let the rated categories be ordered as follow: ‘strongly disagree’, ‘disagree’, ‘neutral’, ‘agree’, and ‘strongly agree’.
The weighted kappa coefficient takes into consideration the different levels of disagreement between categories. For example, if one rater ‘strongly disagrees’ and another ‘strongly agrees’ this must be considered a greater level of disagreement than when one rater ‘agrees’ and another ‘strongly agrees’ (Tang et al. 2015).
Formula
kxk contingency table. Let’s consider the following k×k contingency table summarizing the ratings scores from two raters. k is the number of categories. The table cells contain the counts of cross-classified categories. These counts are indicated by the notation n11, n12, ..., n1K for row 1; n21, n22, ..., n2K for row 2 and so on.
## rater2
## rater1 Level.1 Level.2 Level... Level.k Total
## Level.1 n11 n12 ... n1k n1+
## Level.2 n21 n22 ... n2k n2+
## Level... ... ... ... ... ...
## Level.k nk1 nk2 ... nkk nk+
## Total n+1 n+2 ... n+k NTerminologies:
- The column “Total” (
n1+, n2+, ..., nk+) indicates the sum of each row, known as row margins or marginal counts. Here, the total sum of a given rowiis namedni+. - The row “Total” (
n+1, n+2, ..., n+k) indicates the sum of each column, known as column margins. Here, the total sum of a given columniis namedn+i - N is the total sum of all table cells
- For a give row/column, the marginal proportion is the row/column margin divide by N. This is also known as the marginal frequencies or probabilities. For a row
i, the marginal proportion isPi+ = ni+/N. Similarly, for a given columni, the marginal proportion isP+i = n+i/N. - For each table cell, the proportion can be calculated as the cell count divided by N.
Joint proportions. The proportion in each cell is obtained by dividing the count in the cell by total N cases (sum of the all the table counts).
## rater2
## rater1 Level.1 Level.2 Level... Level.k Total
## Level.1 p11 p12 ... p1k p1+
## Level.2 p21 p22 ... p2k p2+
## Level... ... ... ... ... ...
## Level.k pk1 pk2 ... pkk pk+
## Total p+1 p+2 ... p+k 1Weights. To compute a weighted kappa, weights are assigned to each cell in the contingency table. The weights range from 0 to 1, with weight = 1 assigned to all diagonal cells (corresponding to where both raters agree)(Friendly, Meyer, and Zeileis 2015). The type of commonly used weighting schemes are explained in the next sections.
The proportion of observed agreement (Po) is the sum of weighted proportions.
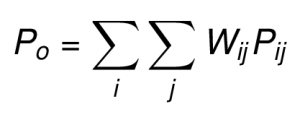
The proportion of expected chance agreement (Pe) is the sum of the weighted product of rows and columns marginal proportions.
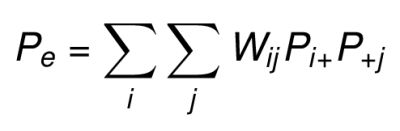
The weighted Kappa can be then calculated by plugging these weighted Po and Pe in the following formula:
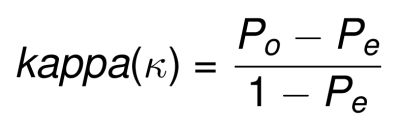
kappa can range form -1 (no agreement) to +1 (perfect agreement).
- when k = 0, the agreement is no better than what would be obtained by chance.
- when k is negative, the agreement is less than the agreement expected by chance.
- when k is positive, the rater agreement exceeds chance agreement.
Note that for 2x2 table (binary rating scales), there is no weighted version of kappa, since kappa remains the same regardless of the weights used.
Types of weights: Linear and quadratic
There are two commonly used weighting system in the literature:
- The Cicchetti-Allison weights (Cicchetti and Allison 1971) based on equal spacing weights for near-match. This also known as linear weights because it is proportional to the deviation of individual rating.
- The Fleiss-Cohen weights (Fleiss and Cohen 1973), based on an inverse-square spacing. This is also known as quadratic weights because it is proportional to the square of the deviation of individual ratings.
For an RxR contingency table,
- the linear weight for a given cell is:
W_ij = 1-(|i-j|)/(R-1) - the quadratic weight for a given cell is:
W_ij = 1-(|i-j|)^2/(R-1)^2
were, |i-j| is the distance between categories and R is the number o categories.
Example of linear weights for a 4x4 table, where two clinical specialist classifies patients into 4 groups:
## Doctor2
## Doctor1 Stade I Stade II Stade III Stade IV
## Stade I 1 2/3 1/3 0
## Stade II 2/3 1 2/3 1/3
## Stade III 1/3 2/3 1 2/3
## Stade IV 0 1/3 2/3 1Example of quadratic weights:
## Doctor2
## Doctor1 Stade I Stade II Stade III Stade IV
## Stade I 1 8/9 5/9 0
## Stade II 8/9 1 8/9 5/9
## Stade III 5/9 8/9 1 8/9
## Stade IV 0 5/9 8/9 1Note that, the quadratic weights attach greater importance to near disagreements. For example, in the situation where you have one category difference between the two doctors diagnosis, the linear weight is 2/3 (0.66). This can be seen as the doctors are in two-thirds agreement (or alternatively, one-third disagreement).
However, the corresponding quadratic weight is 8/9 (0.89), which is strongly higher and gives almost full credit (90%) when there are only one category disagreement between the two doctors in evaluating the disease stage.
However, notice that the quadratic weight drops quickly when there are two or more category differences.
The table below compare the two weighting system side-by-side for 4x4 table:
| Difference | Linear | Quadratic |
|---|---|---|
| 0 | 1 | 1 |
| 1 | 0.66 | 0.89 |
| 2 | 0.33 | 0.55 |
| 3 | 0 | 0 |
How to choose kappa weighting systems
If you consider each category difference as equally important you should choose linear weights (i.e., equal spacing weights).
In other words:
- Use linear weights when the difference between the first and second category has the same importance as a difference between the second and third category, etc.
- Use quadratic weights if the difference between the first and second category is less important than a difference between the second and third category, etc.
Interpretation: Magnitude of the agreement
The interpretation of the magnitude of weighted kappa is like that of unweighted kappa (Joseph L. Fleiss 2003). For most purposes,
- values greater than 0.75 or so may be taken to represent excellent agreement beyond chance,
- values below 0.40 or so may be taken to represent poor agreement beyond chance, and
- values between 0.40 and 0.75 may be taken to represent fair to good agreement beyond chance.
Read more on kappa interpretation at (Chapter @ref(cohen-s-kappa)).
Assumptions
Your data should met the following assumptions for computing weighted kappa.
- You have two outcome categorical variables, which should be ordinal
- The two outcome variables should have exactly the same categories
- You have paired observations; each subject is categorized twice by two independent raters or methods.
- The same two raters are used for all participants.
Statistical hypotheses
- Null hypothesis (H0):
kappa = 0. The agreement is the same as chance agreement. - Alternative hypothesis (Ha):
kappa ≠ 0. The agreement is different from chance agreement.
Example of data
We’ll use the anxiety demo dataset where two clinical doctors classify 50 individuals into 4 ordered anxiety levels: “normal” (no anxiety), “moderate”, “high”, “very high”.
The data is organized in the following 3x3 contingency table:
anxiety <- as.table(
rbind(
c(11, 3, 1, 0), c(1, 9, 0, 1),
c(0, 1, 10, 0 ), c(1, 2, 0, 10)
)
)
dimnames(anxiety) <- list(
Doctor1 = c("Normal", "Moderate", "High", "Very high"),
Doctor2 = c("Normal", "Moderate", "High", "Very high")
)
anxiety## Doctor2
## Doctor1 Normal Moderate High Very high
## Normal 11 3 1 0
## Moderate 1 9 0 1
## High 0 1 10 0
## Very high 1 2 0 10Note that the factor levels must be in the correct order, otherwise the results will be wrong.
Computing Weighted kappa
The R function Kappa() [vcd package] can be used to compute unweighted and weighted Kappa. To specify the type of weighting, use the option weights, which can be either “Equal-Spacing” or “Fleiss-Cohen”.
Note that, the unweighted Kappa represents the standard Cohen’s Kappa which should be considered only for nominal variables. You can read more in the dedicated chapter.
library("vcd")
# Compute kapa
res.k <- Kappa(anxiety)
res.k## value ASE z Pr(>|z|)
## Unweighted 0.733 0.0752 9.75 1.87e-22
## Weighted 0.747 0.0791 9.45 3.41e-21# Confidence intervals
confint(res.k)##
## Kappa lwr upr
## Unweighted 0.586 0.881
## Weighted 0.592 0.903# Summary showing the weights assigned to each cell
summary(res.k)## value ASE z Pr(>|z|)
## Unweighted 0.733 0.0752 9.75 1.87e-22
## Weighted 0.747 0.0791 9.45 3.41e-21
##
## Weights:
## [,1] [,2] [,3] [,4]
## [1,] 1.000 0.667 0.333 0.000
## [2,] 0.667 1.000 0.667 0.333
## [3,] 0.333 0.667 1.000 0.667
## [4,] 0.000 0.333 0.667 1.000Note that, in the above results ASE is the asymptotic standard error of the kappa value.
In our example, the weighted kappa (k) = 0.73, which represents a good strength of agreement (p < 0.0001). In conclusion, there was a statistically significant agreement between the two doctors.
Report
Weighted kappa (kw) with linear weights (Cicchetti and Allison 1971) was computed to assess if there was agreement between two clinical doctors in diagnosing the severity of anxiety. 50 participants were enrolled and were classified by each of the two doctors into 4 ordered anxiety levels: “normal”, “moderate”, “high”, “very high”.
There was a statistically significant agreement between the two doctors, kw = 0.75 (95% CI, 0.59 to 0.90), p < 0.0001. The strength of agreement was classified as good according to Fleiss et al. (2003).
Summary
This chapter explains the basics and the formula of the weighted kappa, which is appropriate to measure the agreement between two raters rating in ordinal scales. We also show how to compute and interpret the kappa values using the R software. Other variants of inter-rater agreement measures are: the Cohen’s Kappa (unweighted) (Chapter @ref(cohen-s-kappa)), which only counts for strict agreement; Fleiss kappa for situations where you have two or more raters (Chapter @ref(fleiss-kappa)).
References
Cicchetti, Domenic V., and Truett Allison. 1971. “A New Procedure for Assessing Reliability of Scoring Eeg Sleep Recordings.” American Journal of EEG Technology 11 (3). Taylor; Francis: 101–10. doi:10.1080/00029238.1971.11080840.
Cohen, J. 1968. “Weighted Kappa: Nominal Scale Agreement with Provision for Scaled Disagreement or Partial Credit.” Psychological Bulletin 70 (4): 213—220. doi:10.1037/h0026256.
Fleiss, Joseph L., and Jacob Cohen. 1973. “The Equivalence of Weighted Kappa and the Intraclass Correlation Coefficient as Measures of Reliability.” Educational and Psychological Measurement 33 (3): 613–19. doi:10.1177/001316447303300309.
Friendly, Michael, D. Meyer, and A. Zeileis. 2015. Discrete Data Analysis with R: Visualization and Modeling Techniques for Categorical and Count Data. 1st ed. Chapman; Hall/CRC.
Joseph L. Fleiss, Myunghee Cho Paik, Bruce Levin. 2003. Statistical Methods for Rates and Proportions. 3rd ed. John Wiley; Sons, Inc.
Tang, Wan, Jun Hu, Hui Zhang, Pan Wu, and Hua He. 2015. “Kappa Coefficient: A Popular Measure of Rater Agreement.” Shanghai Archives of Psychiatry 27 (February): 62–67. doi:10.11919/j.issn.1002-0829.215010.
Warrens, Matthijs J. 2013. “Weighted Kappas for 3x3 Tables.” Journal of Probability and Statistics. doi:https://doi.org/10.1155/2013/325831.
Recommended for you
This section contains best data science and self-development resources to help you on your path.
Books - Data Science
Our Books
- Practical Guide to Cluster Analysis in R by A. Kassambara (Datanovia)
- Practical Guide To Principal Component Methods in R by A. Kassambara (Datanovia)
- Machine Learning Essentials: Practical Guide in R by A. Kassambara (Datanovia)
- R Graphics Essentials for Great Data Visualization by A. Kassambara (Datanovia)
- GGPlot2 Essentials for Great Data Visualization in R by A. Kassambara (Datanovia)
- Network Analysis and Visualization in R by A. Kassambara (Datanovia)
- Practical Statistics in R for Comparing Groups: Numerical Variables by A. Kassambara (Datanovia)
- Inter-Rater Reliability Essentials: Practical Guide in R by A. Kassambara (Datanovia)
Others
- R for Data Science: Import, Tidy, Transform, Visualize, and Model Data by Hadley Wickham & Garrett Grolemund
- Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems by Aurelien Géron
- Practical Statistics for Data Scientists: 50 Essential Concepts by Peter Bruce & Andrew Bruce
- Hands-On Programming with R: Write Your Own Functions And Simulations by Garrett Grolemund & Hadley Wickham
- An Introduction to Statistical Learning: with Applications in R by Gareth James et al.
- Deep Learning with R by François Chollet & J.J. Allaire
- Deep Learning with Python by François Chollet
Version:




No Comments